Section 13.1 Correlation GIFs
Note regression lines and confidence bands can be added using
geom_smooth(method=lm, se=T)
Subsection 13.1.1 N=10, both variables drawn from a uniform distribution
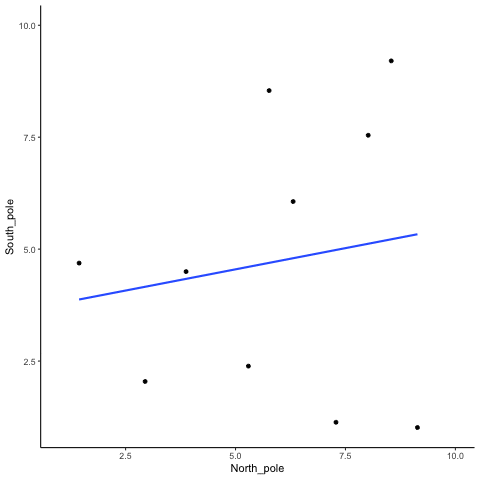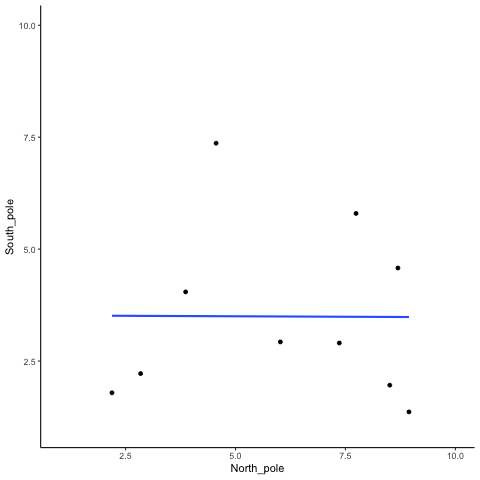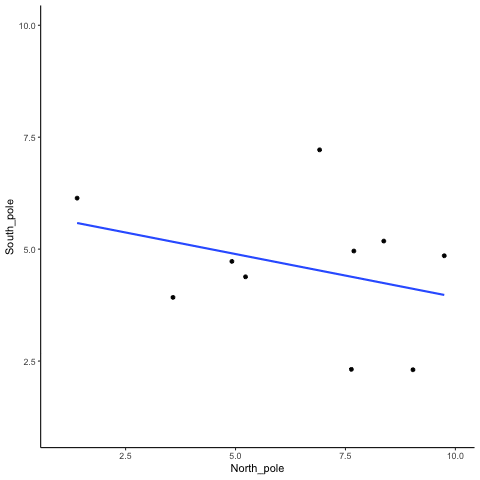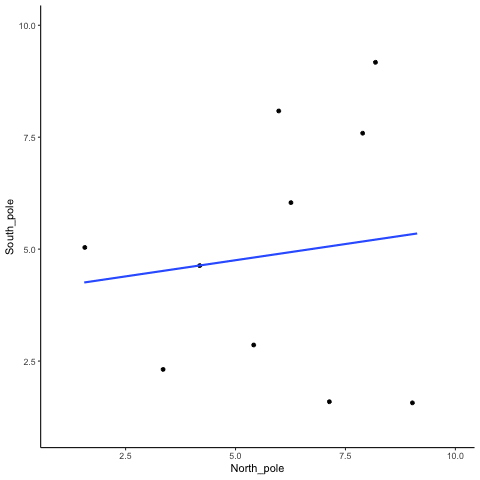
all_df<-data.frame()
for(sim in 1:10){
North_pole <- runif(10,1,10)
South_pole <- runif(10,1,10)
t_df<-data.frame(simulation=rep(sim,10),
North_pole,
South_pole)
all_df<-rbind(all_df,t_df)
}
ggplot(all_df,aes(x=North_pole,y=South_pole))+
geom_point()+
geom_smooth(method=lm, se=FALSE)+
theme_classic()+
transition_states(
simulation,
transition_length = 2,
state_length = 1
)+enter_fade() +
exit_shrink() +
ease_aes('sine-in-out')
Subsection 13.1.2 Correlation between random deviates from uniform distribution across four sample sizes
N= 10,50,100,1000 All values sampled from a uniform distribution
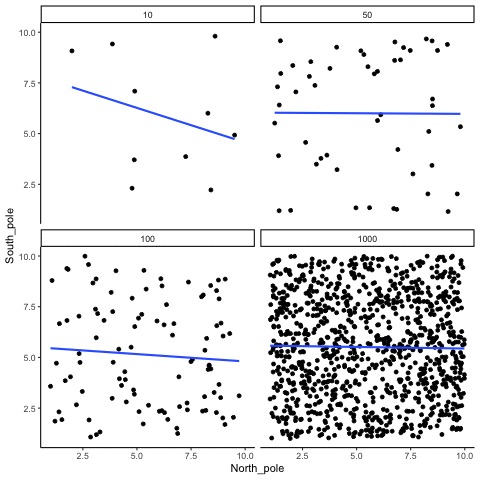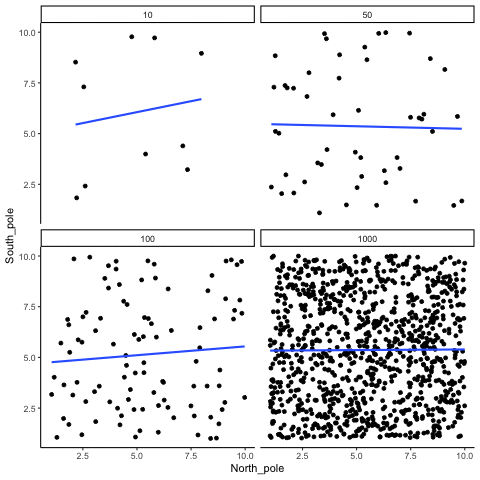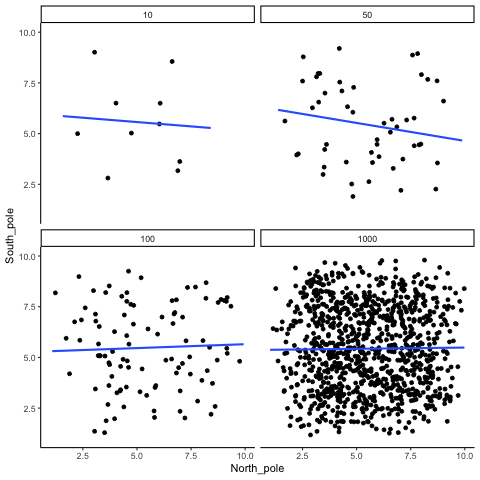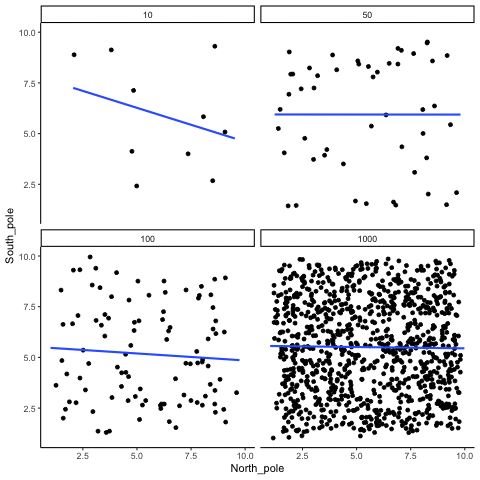
all_df<-data.frame()
for(sim in 1:10){
for(n in c(10,50,100,1000)){
North_pole <- runif(n,1,10)
South_pole <- runif(n,1,10)
t_df<-data.frame(nsize=rep(n,n),
simulation=rep(sim,n),
North_pole,
South_pole)
all_df<-rbind(all_df,t_df)
}
}
ggplot(all_df,aes(x=North_pole,y=South_pole))+
geom_point()+
geom_smooth(method=lm, se=FALSE)+
theme_classic()+
facet_wrap(~nsize)+
transition_states(
simulation,
transition_length = 2,
state_length = 1
)+enter_fade() +
exit_shrink() +
ease_aes('sine-in-out')
Subsection 13.1.3 Correlation between random deviates from normal distribution across four sample sizes
N= 10,50,100,1000 All values sampled from the same normal distribution (mean=0, sd=1)
all_df<-data.frame()
for(sim in 1:10){
for(n in c(10,50,100,1000)){
North_pole <- rnorm(n,0,1)
South_pole <- rnorm(n,0,1)
t_df<-data.frame(nsize=rep(n,n),
simulation=rep(sim,n),
North_pole,
South_pole)
all_df<-rbind(all_df,t_df)
}
}
ggplot(all_df,aes(x=North_pole,y=South_pole))+
geom_point()+
geom_smooth(method=lm, se=FALSE)+
theme_classic()+
facet_wrap(~nsize)+
transition_states(
simulation,
transition_length = 2,
state_length = 1
)+enter_fade() +
exit_shrink() +
ease_aes('sine-in-out')
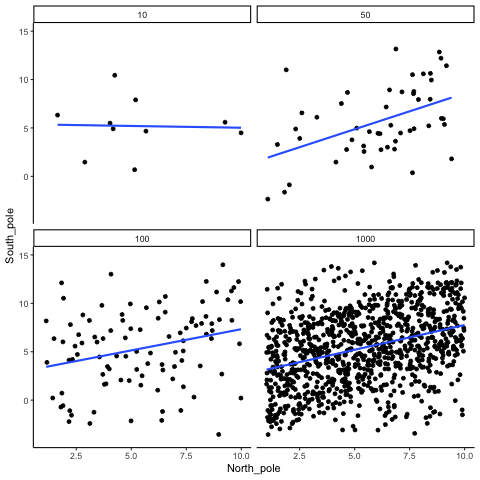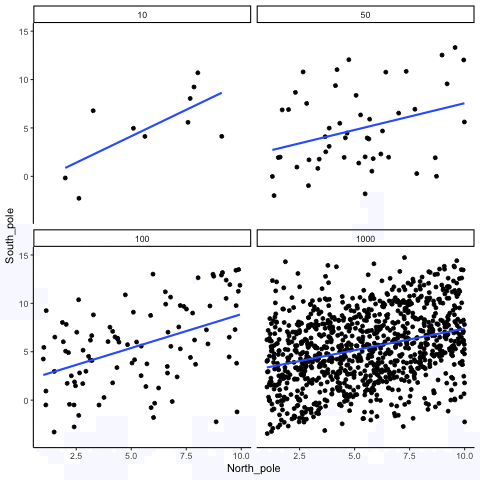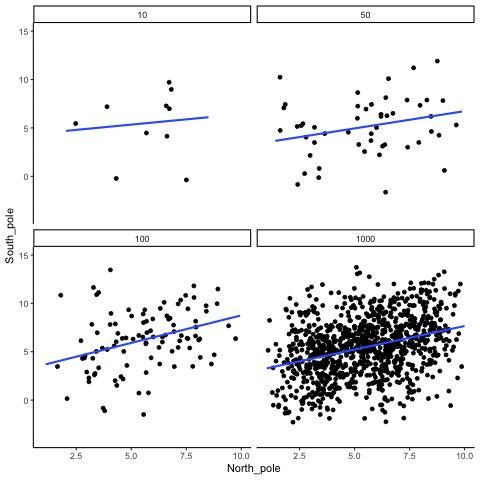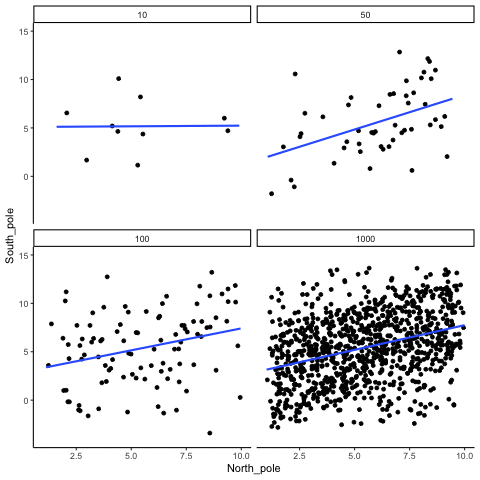
Subsection 13.1.4 Correlation between X and Y variables that have a true correlation as a function of sample-size
Sample-sizes = 10, 50, 100, 1000. There is a true correlation between X and Y in this case, set at r=.5.
library(MASS)
all_df<-data.frame()
for(sim in 1:10){
for(n in c(10,50,100,1000)){
d <- mvrnorm(n = n, mu = c(0,0), Sigma = matrix(c(1,.5,.5,1), ncol = 2),
empirical = FALSE)
t_df<-data.frame(nsize=rep(n,n),
simulation=rep(sim,n),
North_pole=d[,1],
South_pole=d[,2])
all_df<-rbind(all_df,t_df)
}
}
ggplot(all_df,aes(x=North_pole,y=South_pole))+
geom_point()+
geom_smooth(method=lm, se=FALSE)+
theme_classic()+
facet_wrap(~nsize)+
transition_states(
simulation,
transition_length = 2,
state_length = 1
)+enter_fade() +
exit_shrink() +
ease_aes('sine-in-out')
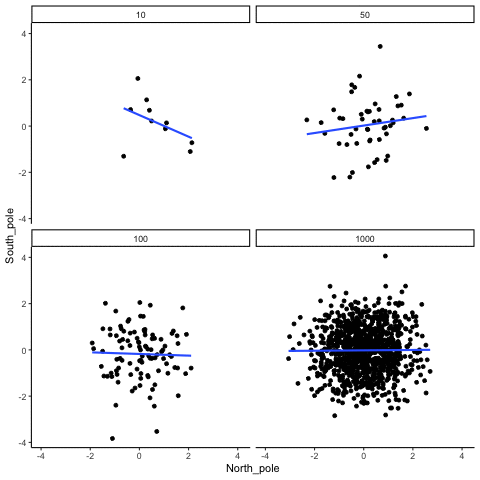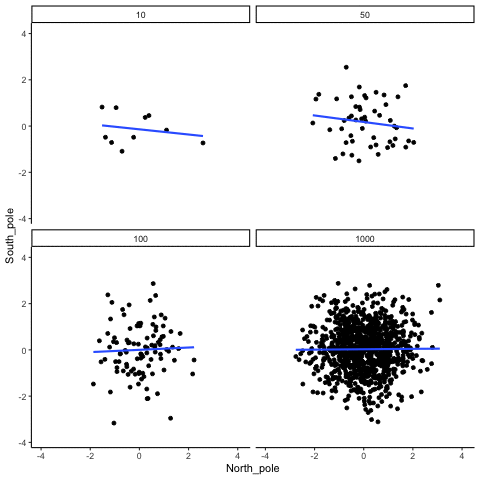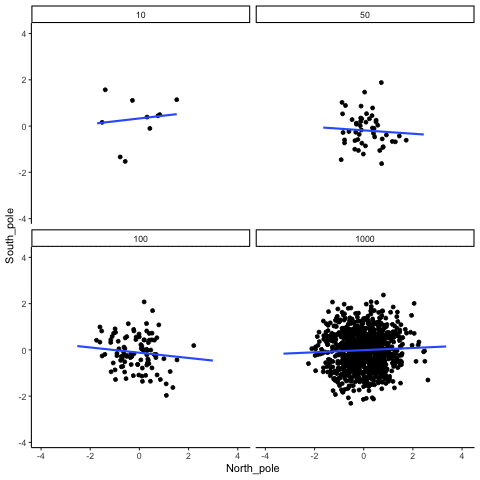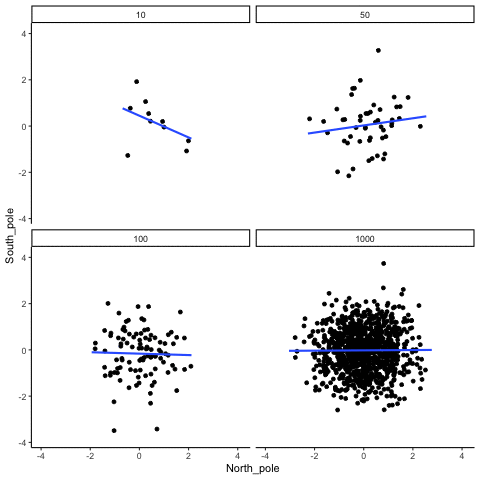
Subsection 13.1.5 Type I errors, sampling random deviates from normal distribution with regression lines
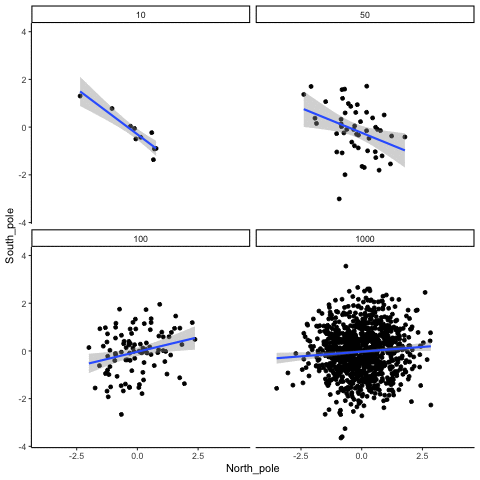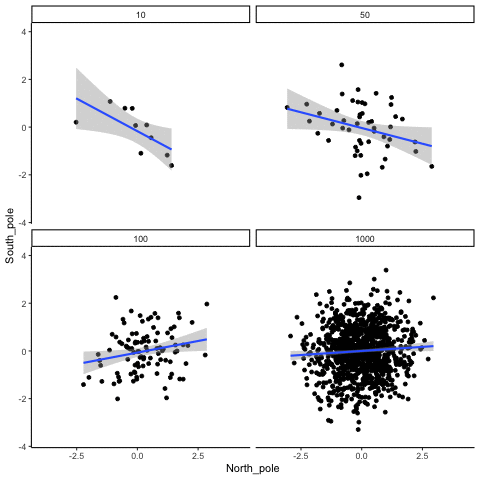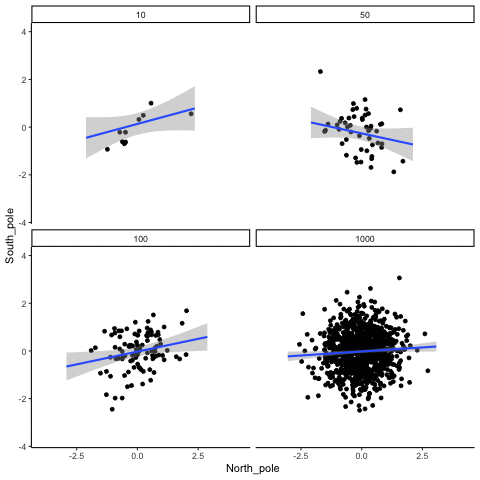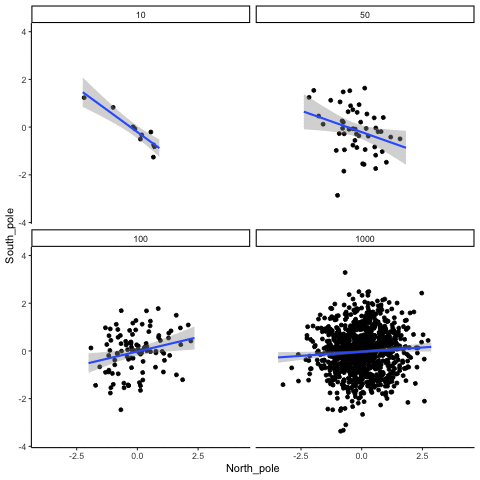
Four sample-sizes, N= 10, 50, 100, 1000. All samples drawn from normal distribution (u=0, sd=1). Shows only the samples where p<.05. This is only true for about 5% of all samples (shown here for first 10 Type I errors for each N). Thus, we see Type I errors (significant correlations when X and Y are both completely random).
all_df<-data.frame()
for(n in c(10,50,100,1000)){
count_sims<-0
for(sim in 1:1000){
North_pole <- rnorm(n,0,1)
South_pole <- rnorm(n,0,1)
if(cor.test(North_pole,South_pole)$p.value<.05){
count_sims<-count_sims+1
t_df<-data.frame(nsize=rep(n,n),
simulation=rep(count_sims,n),
North_pole,
South_pole)
all_df<-rbind(all_df,t_df)
if(count_sims==10){
break
}
}
}
}
ggplot(all_df,aes(x=North_pole,y=South_pole))+
geom_point()+
geom_smooth(method=lm, se=TRUE)+
theme_classic()+
facet_wrap(~nsize)+
transition_states(
simulation,
transition_length = 2,
state_length = 1
)+enter_fade() +
exit_shrink() +
ease_aes('sine-in-out')
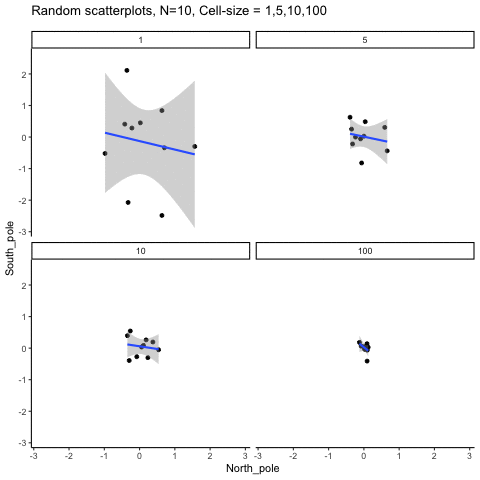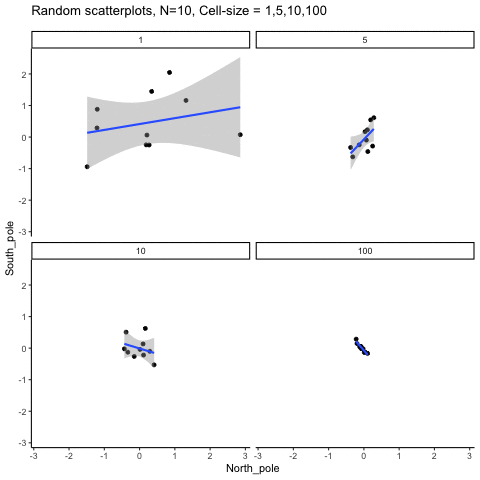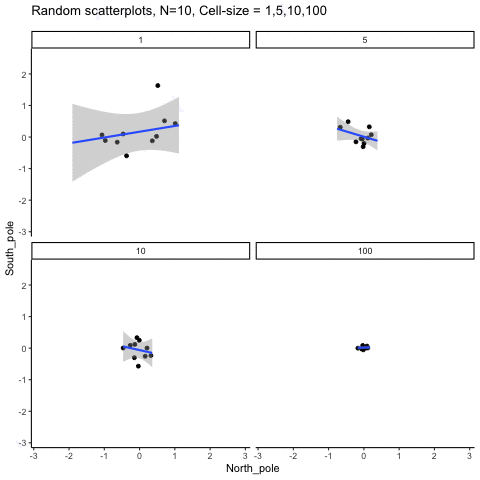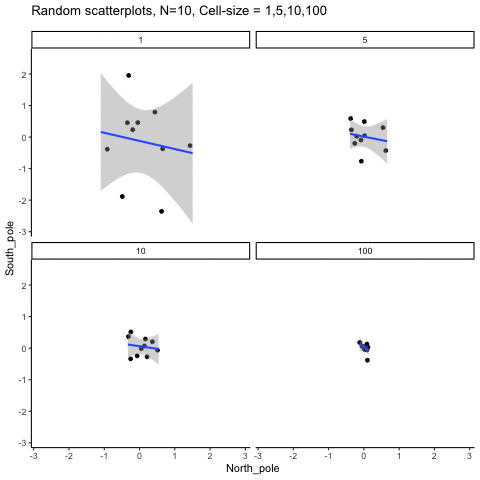
Subsection 13.1.6 Cell-size and correlation
This simulation illustrates how the behavior of correlating two random normal samples as a function of cell-size. The sample-size is always set at N=10. For each panel, the simulation uses an increasing cell-size to estimate the mean for X and Y. When cell-size is 1, 10 X and Y values are drawn from the same normal (u=0, sd=1). When cell-size is 5, for each X,Y score in the plot, 5 samples were drawn from the same normal, and then the mean of the samples is plotted. The effect of cell-size shrinks the dot cloud, as both X and Y scores provide better estimates of the population mean = 0. Cell-size has no effect on the behavior of r, which swings around because sample-size N is small. These are all random, so there is always a 5% type I error rate (alpha =.05).
get_sampling_means<-function(m,sd,cell_size,s_size){
save_means<-length(s_size)
for(i in 1:s_size){
save_means[i]<-mean(rnorm(cell_size,m,sd))
}
return(save_means)
}
all_df<-data.frame()
for(n in c(1,5,10,100)){
count_sims<-0
for(sim in 1:10){
North_pole <- get_sampling_means(0,1,n,10)
South_pole <- get_sampling_means(0,1,n,10)
count_sims<-count_sims+1
t_df<-data.frame(nsize=rep(n,10),
simulation=rep(count_sims,10),
North_pole,
South_pole)
all_df<-rbind(all_df,t_df)
}
}
ggplot(all_df,aes(x=North_pole,y=South_pole))+
geom_point()+
geom_smooth(method=lm, se=TRUE)+
theme_classic()+
facet_wrap(~nsize)+
ggtitle("Random scatterplots, N=10, Cell-size = 1,5,10,100")+
transition_states(
simulation,
transition_length = 2,
state_length = 1
)+enter_fade() +
exit_shrink() +
ease_aes('sine-in-out')
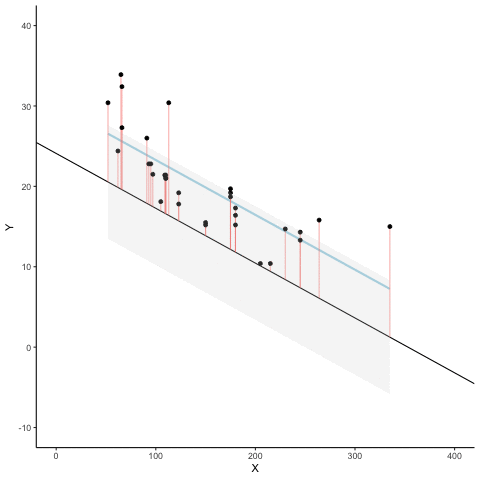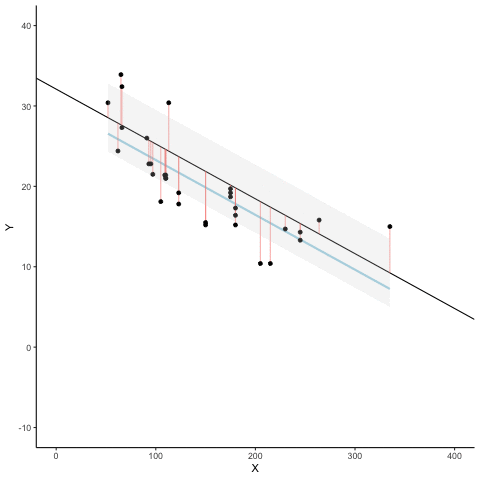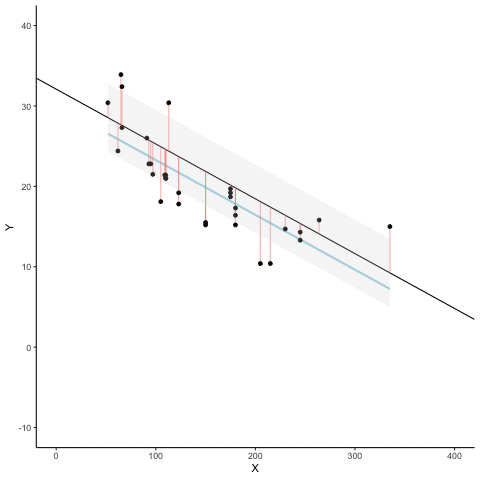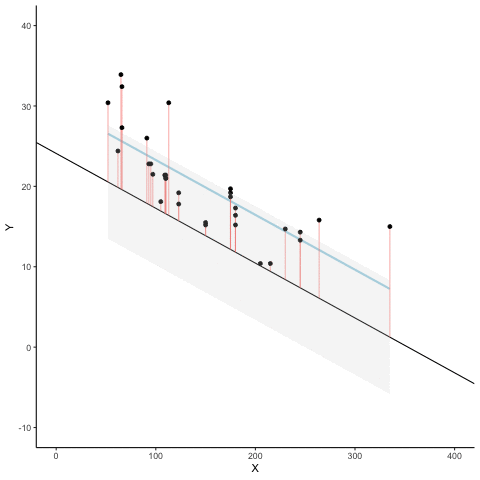
Subsection 13.1.7 Regression
We look at how the residuals (error from points to line) behave as the regression lines moves above and below it’s true value. The total error associated with all of the red lines is represents by the grey area. This total error is smallest (minimized) when the black line overlaps with the blue regression line (the best fit line). The total error expands as the black line moves away from the regression. That’s why the regression line is the least wrong (best fit) line to skewer the data (according to least squares definition)
d <- mtcars
fit <- lm(mpg ~ hp, data = d)
d$predicted <- predict(fit) # Save the predicted values
d$residuals <- residuals(fit) # Save the residual values
coefs<-coef(lm(mpg ~ hp, data = mtcars))
coefs[1]
coefs[2]
x<-d$hp
move_line<-c(seq(-6,6,.5),seq(6,-6,-.5))
total_error<-length(length(move_line))
cnt<-0
for(i in move_line){
cnt<-cnt+1
predicted_y <- coefs[2]*x + coefs[1]+i
error_y <- (predicted_y-d$mpg)^2
total_error[cnt]<-sqrt(sum(error_y)/32)
}
move_line_sims<-rep(move_line,each=32)
total_error_sims<-rep(total_error,each=32)
sims<-rep(1:50,each=32)
d<-d %>% slice(rep(row_number(), 50))
d<-cbind(d,sims,move_line_sims,total_error_sims)
anim<-ggplot(d, aes(x = hp, y = mpg, frame=sims)) +
geom_smooth(method = "lm", se = FALSE, color = "lightblue") +
geom_abline(intercept = 30.09886+move_line_sims, slope = -0.06822828)+
lims(x = c(0,400), y = c(-10,40))+
geom_segment(aes(xend = hp, yend = predicted+move_line_sims, color="red"), alpha = .5) +
geom_point() +
geom_ribbon(aes(ymin = predicted+move_line_sims - total_error_sims, ymax = predicted+move_line_sims + total_error_sims), fill = "lightgrey", alpha=.2)+
theme_classic()+
theme(legend.position="none")+
xlab("X")+ylab("Y")+
transition_manual(frames=sims)+
enter_fade() +
exit_fade()+
ease_aes('sine-in-out')
animate(anim,fps=5)

